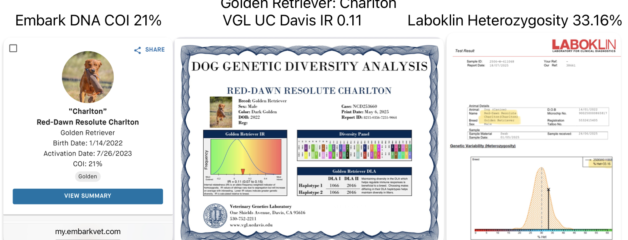
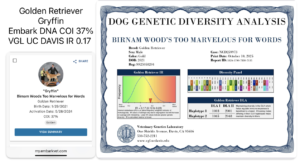
UC Davis VGL vs Embark DNA testing of genetic diversity in dogs
Embark measures a dog’s genetic inbreeding using DNA from across the whole genome. Multiple scientific studies have shown this method to be an accurate way to estimate true inbreeding in dogs.
On Embark, there are over 8,000 Golden Retrievers with publicly visible results (these are breeder accounts that chose full public access). Embark’s internal database includes many more Goldens that are not public and are used for population-level analysis and private pairing tools.
By comparison, the UC Davis Veterinary Genetics Laboratory (VGL) database includes about 1,200 Golden Retrievers in total.
Comparing how related two dogs are:
💙 VGL compares dogs only within its smaller Golden Retriever dataset (about 1,200 dogs).
❤️ Embark compares dogs against thousands of Goldens, plus an even larger private database, giving a broader picture of how related two dogs really are.
Strengths of each system:
💙 VGL is very useful for comparing haplotypes in each purebred family of dogs. This helps breeders avoid close inbreeding within a family line. However, because the dataset is smaller, introducing a new line can sometimes appear to add more diversity than it truly does.
❤️ Embark can compare any two dogs (same breed or different breeds) and estimate the overall genetic inbreeding of potential offspring. It does this by measuring runs of homozygosity across the genome, which is the most accurate way to assess true inbreeding.
About Embark’s COI:
Embark’s COI closely reflects a dog’s real genetic inbreeding. Only whole genome analysis of homozygosity can estimate this accurately. Because Embark is conservative in how it reports this data, its COI values may be slightly lower than a full research grade whole genome profile, but they are still very close to biological reality.