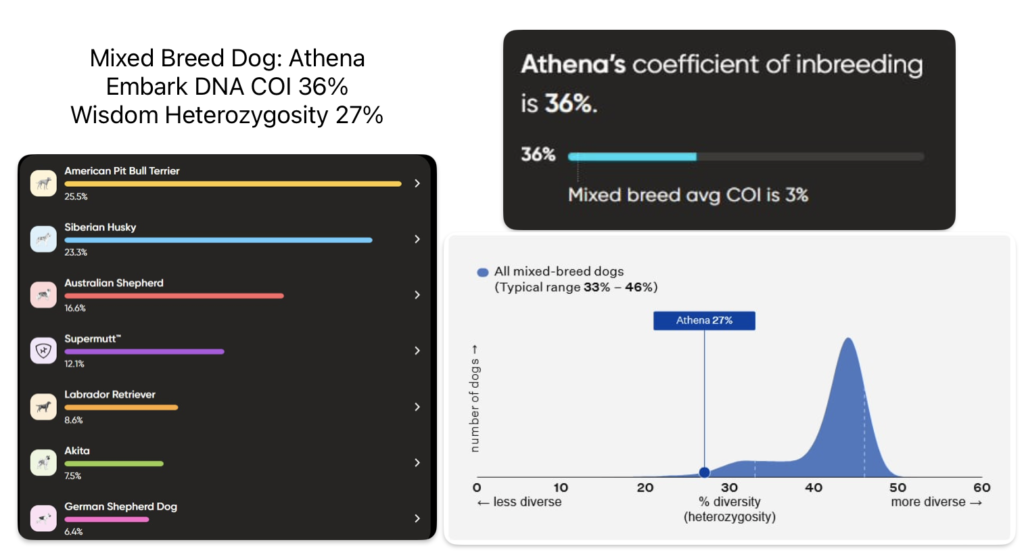
Genetic Diversity is shown here in picture form of multiple dogs from a variety of breeds and mixes. For each dog at least two platforms of DNA testing of genetic diversity was used. This data helps illustrate easy to understand examples of how each DNA company differs and relates to each other when it comes to the DNA profiling of inbreeding of dogs.
Two types of technology can be used. SNP and STR
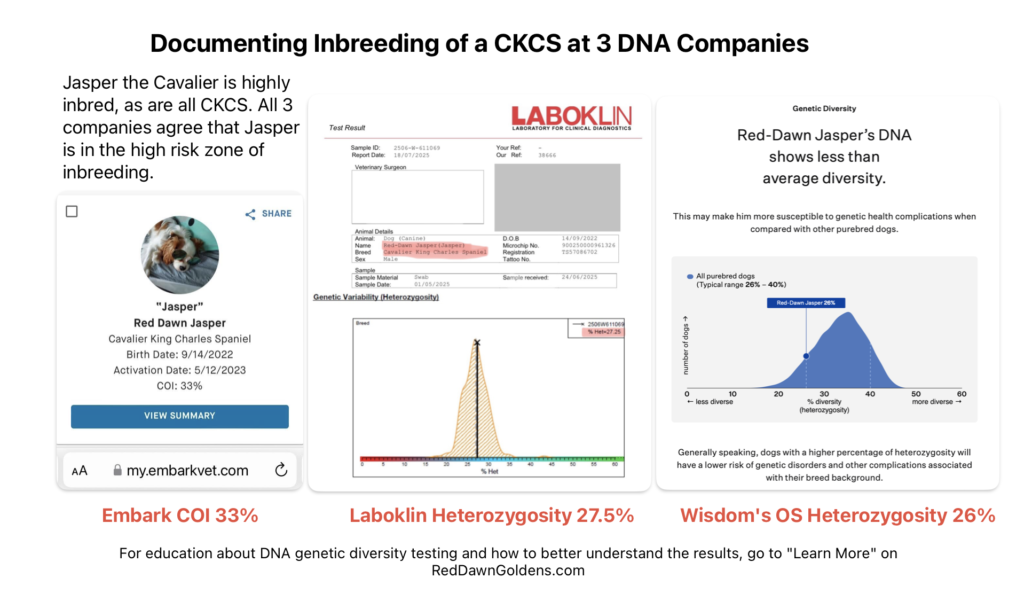
- Canine SNP (Single Nucleotide Polymorphism) testing analyzes many thousands of tiny genetic variations across a dog’s genome to determine breed ancestry, health risks, and traits with high precision. Companies like Embark, Wisdom Panel, Feragen, and Laboklin use this technology to map DNA against large databases.
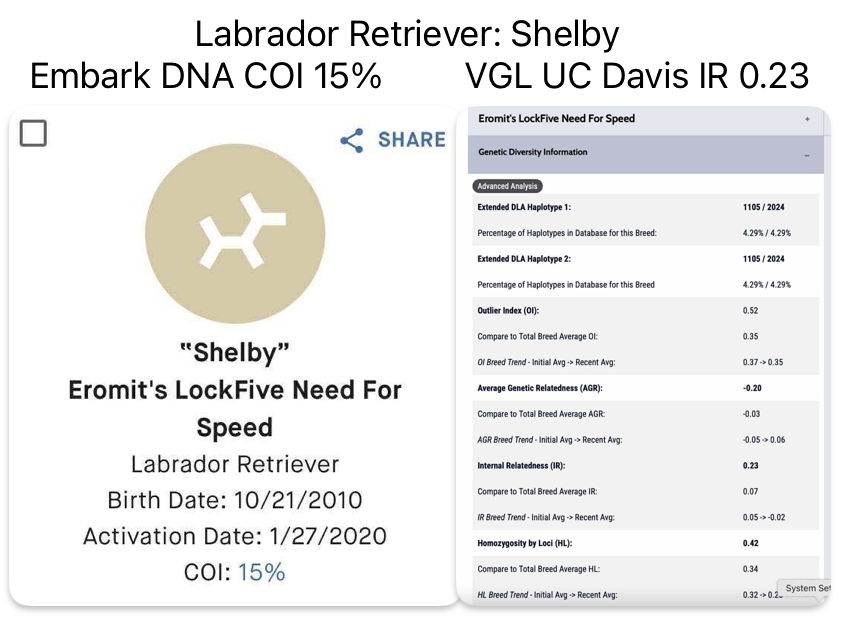
- Canine STR at VGL UC Davis uses 33 Autosomal STR Loci markers located across the genome to measure genetic heterogeneity and diversity.
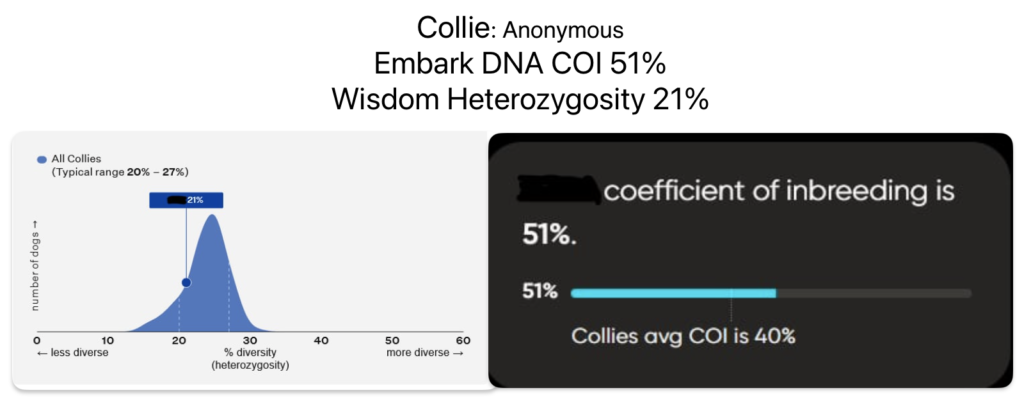
Wisdom Panel’s Optimal Selection, Feragen and Laboklin test a dog’s genetic Heterozygosity by evaluating the DNA. A higher score is better, like in Basketball. A lower score means more inbreeding is found in that dog.
Heterozygosity of 45% is VERY GOOD
Heterozygosity of 15% is VERY BAD
Embark DNA tests a dog’s inbreeding by looking at runs of homozygosity, called COI. A lower score is better, like in Golf. A higher score means more inbreeding is found in that dog. COI is the abbreviation for Coefficient of Inbreeding.
COI of 0-10% is VERY GOOD
COI of 35%+ is VERY BAD
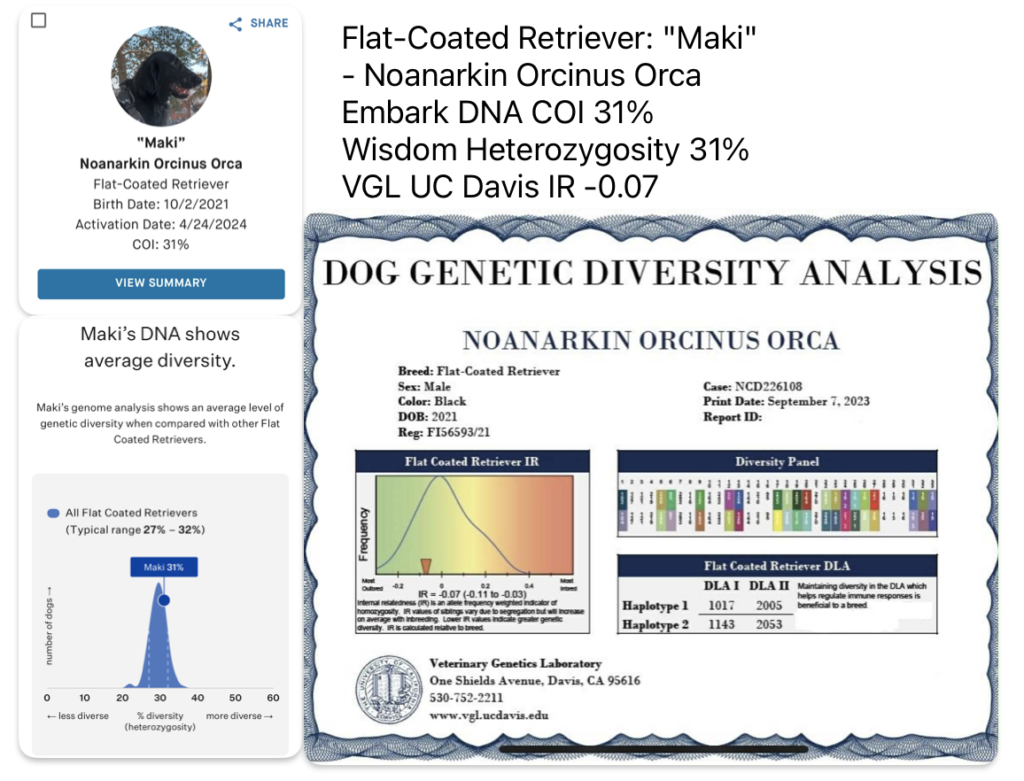
VGL UC Davis test shows IR, Internal Relatedness by evaluating homozygosity. As you can see in the colored graph, the lower the score the better (in the Green portion) and the higher the score the worse (in the Red portion). Think of IR as how Golf is scored, the low score is best.
To easily remember the differences between the platforms of DNA testing of genetic diversity:
COI = Golf, the lower the score the better (winner)
IR = Golf, the lower the score the better (winner)
Heterozygosity = Basketball, the higher the score the better (winner)
Pictured are multiple dogs with at least two platform’s DNA tests for genetic diversity.
Want to see only Golden Retrievers tested for genetic diversity? Check out the Golden’s Diversity page
*** The owners of these dogs graciously submitted their dog’s testing results to Alicia Dawn of Red-Dawn Goldens and Cavaliers. These dog’s owners are helping the public by allowing their dog’s data help educate all of us. We are grateful to each dog’s owner for their contribution to the education of understanding inbreeding in dogs.

Una is a mixed breed dog showing high heterozygosity and very low inbreeding from DNA testing at Embark and Wisdom Panel.
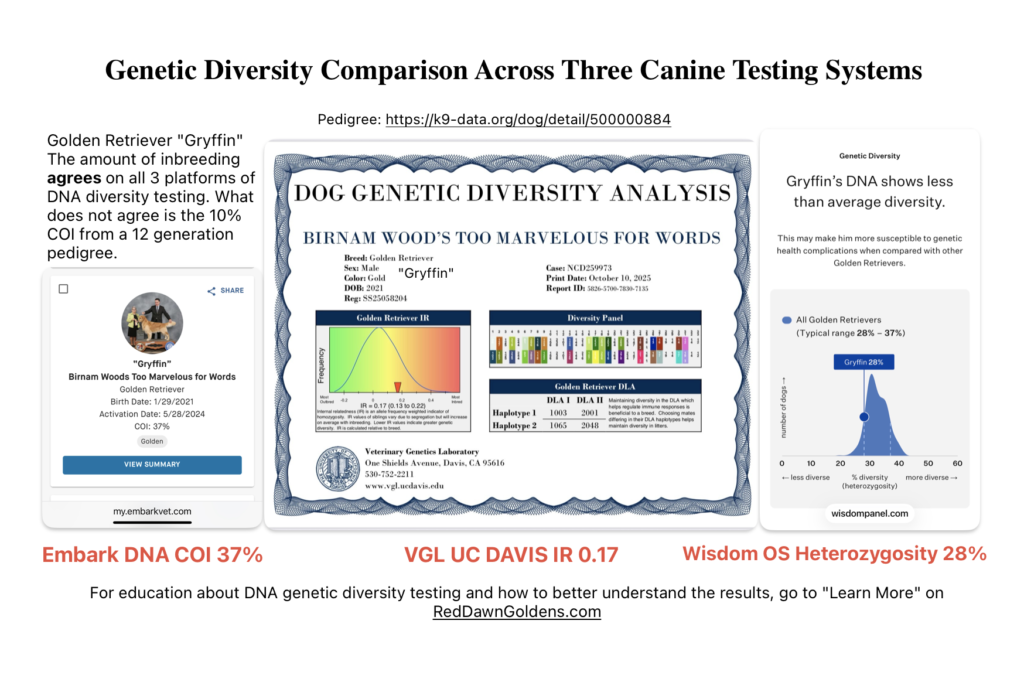
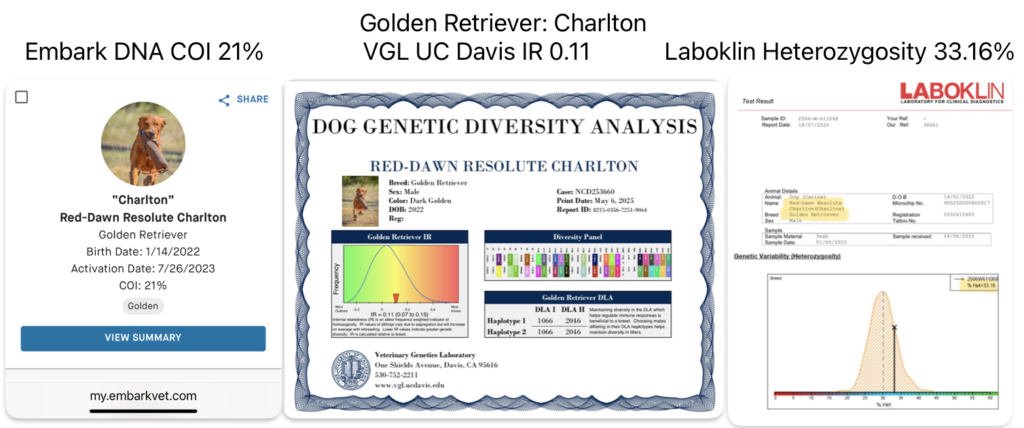
DNA genetic diversity testing of a male Golden Retriever at VGL, Embark and Wisdom Panel Optimal Selection. He is considered moderate to highly inbred on all three systems.

Both these Border Collies are low on their levels of inbreeding as scored by two DNA companies, Embark and Wisdom Panel
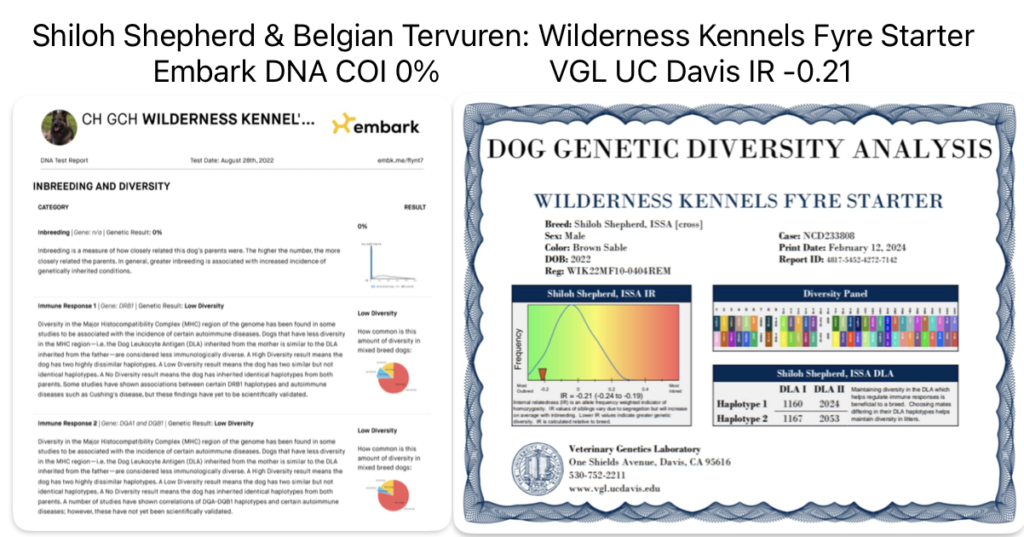
This dog who is part Shiloh Shepherd is incredibly genetically diverse and is not inbred based on two companies DNA testing, Embark and UC Davis VGL
References
Zimmerman SJ, Aldridge CL, Oyler-McCance SJ. An empirical comparison of population genetic analyses using microsatellite and SNP data for a species of conservation concern. BMC Genomics. 2020 Jun 1;21(1):382. doi: 10.1186/s12864-020-06783-9. PMID: 32487020; PMCID: PMC7268520.
https://pubmed.ncbi.nlm.nih.gov/32487020/
This study shows that SNPs have three main advantages over microsatellites (STRs): they give more accurate measures of genetic diversity, better identify groups within a population, and can detect local adaptation. Overall, SNPs are becoming more useful for studying genetic diversity, especially in conservation.
Pérez-González, J.; Carranza, J.; Anaya, G.; Broggini, C.; Vedel, G.; de la Peña, E.; Membrillo, A. Comparative Analysis of Microsatellite and SNP Markers for Genetic Management of Red Deer. Animals 2023, 13, 3374. https://doi.org/10.3390/ani13213374
https://www.mdpi.com/2076-2615/13/21/3374
Both microsatellites (STRs) and SNPs gave similar overall results for population structure and genetic diversity. STRs were useful for getting a general picture of the population. However, SNPs were more precise, better at detecting subtle patterns, and could find relationships (like inbreeding effects) that STRs missed. Because of this, researchers recommend using SNPs when possible.
Vignal A, Milan D, SanCristobal M, Eggen A. A review on SNP and other types of molecular markers and their use in animal genetics. Genet Sel Evol. 2002 May-Jun;34(3):275-305. doi: 10.1186/1297-9686-34-3-275. PMID: 12081799; PMCID: PMC2705447.
https://link.springer.com/article/10.1186/1297-9686-34-3-275
DNA markers have become more important in animal genetics. Microsatellites (STRs) were widely used because they are easy to test and provide a lot of information. However, SNPs, despite having only two variants, have become more popular. This review explains why SNPs are now preferred and how they compare to other marker types.
Liu N, Chen L, Wang S, Oh C, Zhao H. Comparison of single-nucleotide polymorphisms and microsatellites in inference of population structure. BMC Genet. 2005 Dec 30;6 Suppl 1(Suppl 1):S26. doi: 10.1186/1471-2156-6-S1-S26. PMID: 16451635; PMCID: PMC1866760.
https://pmc.ncbi.nlm.nih.gov/articles/PMC1866760/
SNPs are widely used to study genetics, but they are often compared to microsatellites (STRs). In this study of 236 people, researchers analyzed 328 STRs and 15,840 SNPs. Each STR was about 4–12 times more informative than a single SNP. However, because there were so many more SNPs, they provided better overall results for understanding population structure.